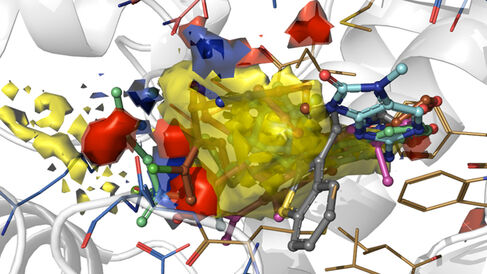
The Blundell Group and our Cryo-Electron Microscopy Facility have published in Nature an exciting breakthrough in understanding how cancer drugs targeting the DNA repair protein DNA-PKcs bind to their target.
Protein kinases are regulators of almost all aspects of cell life and play central roles in signaling pathways. Since the FDA approval of one of the first kinase inhibitors (imatinib) in 2001, targeting protein kinases has shown great potential in drug discovery and impact in the treatment of many diseases, especially cancer.
A hallmark of cancer is genome instability – a high frequency of mutations and damage in a cell's DNA. Cancer cells have higher levels of DNA damage than normal cells and are deficient in one or more DNA damage response and repair pathways that would normally correct such errors. As a result of this, inhibiting a specific human DNA damage response and repair pathway can lead to synthetic lethality and the death of cancer cells. It can also improve the effect of the traditional DNA-damaging and genotoxic treatments of radiotherapy and chemotherapy.
DNA-PKcs (DNA-dependent protein kinase catalytic subunit) is a key kinase regulating human DNA damage response and repair and is indispensable in non-homologous end joining (NHEJ), which is the preferred pathway for the repair of the most toxic DNA double-strand breaks in human genes. DNA-PKcs selective inhibitors, some of which are under phase I and phase II clinical trials, have exhibited high potential as treatment for various cancers. However, due to various limiting factors including the kinase's large size and flexibility, the mode of action of its ligands (including inhibitors and drug candidates) remained unclear, posing a major hurdle to drug development.
In their new publication, the Blundell Group use cryo-electron microscopy (cryo-EM) to reveal how existing small molecules, including drug candidates under clinical trial, interact with DNA-PKcs. Thanks to the revolutionary improvement of single-particle analysis in cryo-EM, the authors unveil how the natural ligand analogue (ATPγS) and different inhibitors (including the broad-spectrum inhibitor wortmannin, the older-generation inhibitor NU7441, and the newer-generation inhibitors AZD7648 and M3814) interact with DNA-PKcs at the molecular level. Moreover, comparisons among the modes of action of natural ligand analogues, synthetic inhibitors with different specificities, and homologous protein-ligand complexes give clues concerning the origin of drug specificity, providing structural insights and ideas for next-generation DNA-PKcs inhibitor development. In addition, the structures reveal the concerted movement caused by ligand binding in the kinase domain, which is important for kinase catalysis.
Now that the structural binding of the small molecules is better understood, there is a clear opportunity for better drug development. Meanwhile, knowledge of the interaction mechanisms of different small molecules, and comparisons to other similar protein-ligand interactions, provide insights on the origin of drug specificity that can guide the future development of more specific small-molecule drugs to this very large kinase.
In general, the publication describes ligand regulation of DNA-PKcs for the first time. The high-resolution structures will allow improved structure-based drug development in the future and demonstrate the huge potential of cryo-EM in structure-guided drug development for other large and challenging targets.
Liang S, Thomas SE, Chaplin AK, Hardwick SW, Chirgadze DY, Blundell TL (2022). Structural insights into inhibitor regulation of the DNA repair protein DNA-PKcs. Nature. doi: 10.1038/s41586-021-04274-9
