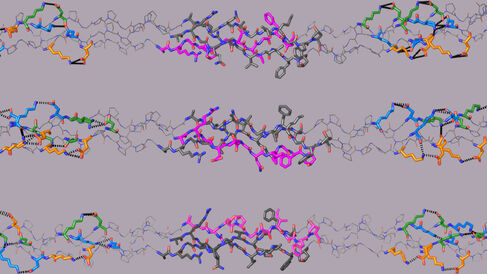
The Farndale Group and their collaborators have published a new paper in Nature Chemical Biology that reveals the molecular assembly of type I collagen.
Collagens are the most abundant proteins present in mammalian skin and bone, and are the structural components of connective tissues. There are 28 known types of collagen which form sponge-like networks or rope-like fibres. Cells embed within and communicate with these dense matrices to multiply and to sense their extracellular environment. Collagens also play prominent roles in cell signalling, blood clotting, cancer progression and cardiovascular disease, and are responsible for genetically inherited diseases such as brittle bone disease and Ehlers-Danlos syndrome.
The large size and highly complex structure of collagens makes them difficult to study on a molecular scale. So much so that the molecular assembly of the most abundant collagen, type I collagen (COL1), has remained an unresolved mystery for nearly five decades.
COL1 forms a triple helix composed of two similar (chains A) and one unique (chain B) polypeptide chains. These chains self-assemble with a one amino acid stagger between them to optimise molecular packing. Consequently, chain B in COL1 could begin in the leading (BAA), middle (ABA) or trailing (AAB) position, resulting in three isomeric biomacromolecules that are expected to differ significantly in their biological activities. The correct positioning of chain B within the native COL1 is unknown, and all three positions have variously been reported over the last 50 years.
In their new paper, the Farndale Group computationally designed three small, triple-helical collagen peptides that mimic the three possible COL1 molecular structures. Only one of these could successfully interact with non-collagenous proteins known to bind to COL1. The AAB mimic demonstrated a clear and strong preference for the receptor tyrosine kinase Discoidin Domain Receptor 1 (DDR1) and the blood serum protein von Willebrand factor (VWF) in solid-phase binding assays, and also induced distinctly higher levels of cellular DDR1 and DDR2 activation compared with the BAA and ABA mimics. These results provide an empirical demonstration that chain B in COL1 resides in the trailing (AAB) position.
Therapeutic agents that disrupt DDR-collagen and VWF-collagen interactions are potential drug targets in preventing cancer progression and hemostasis, respectively. Moreover, numerous point mutations in COL1 cause genetic diseases such as osteogenesis imperfecta, or brittle bone disease. Structural perturbations in COL1 due to mutations in either or both chains can be correctly modelled and correlated with the resulting phenotypic severity only if the alignment of chain B is precisely known. Thus, this new work can potentially advance understanding of both physiological and genetic aspects of collagen function.
